About CBMAP
Project Overview
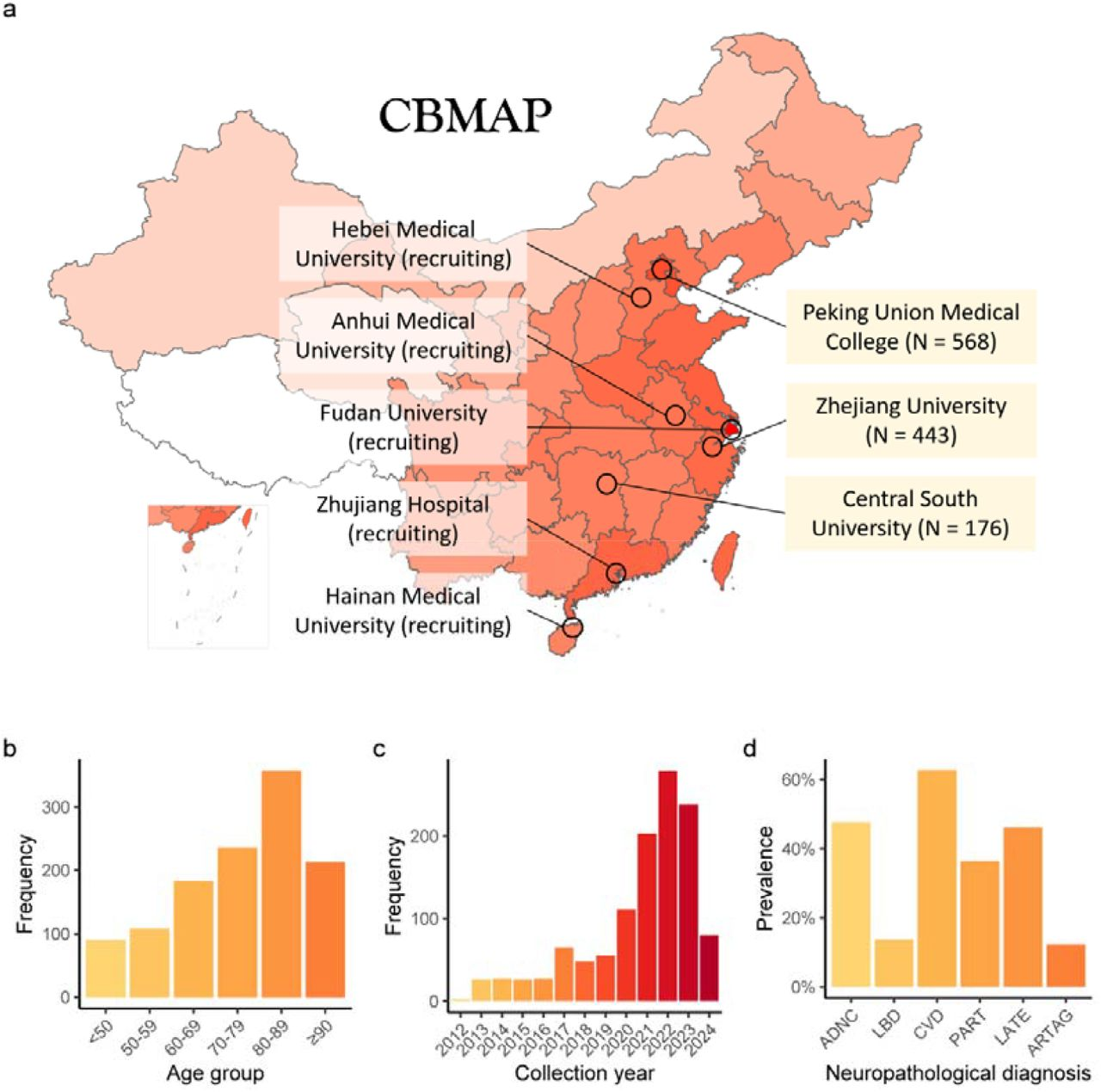
The China Brain Multi-omics Atlas Project (CBMAP) aims to generate a comprehensive molecular reference map of over 1,000 human brains (Phase I), spanning a broad age range and multiple regions in China, to address the underrepresentation of East Asian populations in brain research.
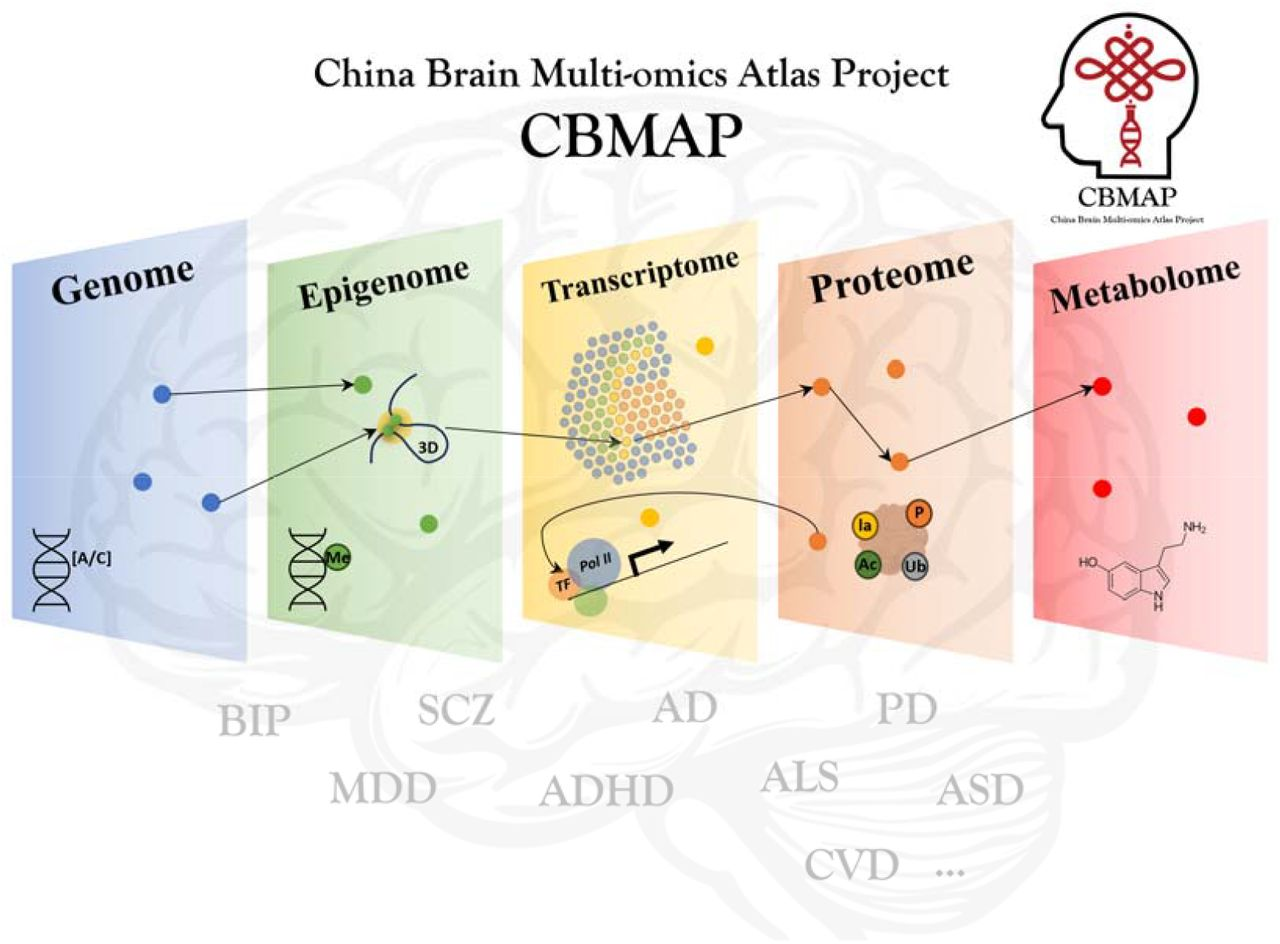
By integrating genome, epigenome, transcriptome, proteome (including multiple post-translational modifications), and metabolome data, CBMAP provides a rich and invaluable resource for investigating the molecular underpinnings of aging-related brain phenotypes and neuropsychiatric disorders.
Leveraging high-throughput omics data and advanced technologies, such as spatial transcriptomics, proteomics, and single-nucleus 3D chromatin structure analysis, this atlas will serve as a crucial resource for the brain science community, illuminating disease mechanisms and enhancing the utility of data from genome-wide association studies (GWAS). CBMAP is also poised to accelerate drug discovery and precision medicine for brain disorders.
Research Goals
- (1) Characterize molecular elements in Chinese population and highlight differences within Chinese and between Chinese and other populations.
- (2) Establish comprehensive atlas of molecules and co-regulation modules associated with aging and neuropsychiatric disorders across multiple layers.
- (3) Generate integrated maps of spatial omics, single-nucleus 3D epigenetic interactions, and transcriptional regulation for representative samples.
- (4) Profile multiple PTMs, uncovering protein-level crosstalks at the population level and identifying PTM quantitative trait loci (ptmQTLs).
- (5) Elucidate genetic architecture of various molecules with ancestry-specific, disease-specific, and age-specific characteristics.
- (6) Enhance mapping resolution in disease-associated molecules by integrating multi-ancestry data under the hypothesis of "different ancestries, same pathogenic genetic element(s)".
Publication
Project Impact
Advancing brain research through comprehensive molecular analysis